Advanced Next-Generation Workflows in PubChemR
Selcuk Korkmaz, Bilge Eren Yamasan, Dincer Goksuluk
2026-03-07
This vignette has two layers.
The first layer is researcher-facing: a concrete compound-triage workflow that answers the question “what can I actually do with PubChemR?”.
The second layer is systems-facing: the transport, caching, async, batching, and typed-result machinery that makes those workflows reproducible and robust.
It is designed to be:
- Offline-first and reproducible by default.
- Structured from basic to advanced usage.
- Practical for production pipelines.
- Explicit about object contracts, failures, and edge cases.
How to use this guide:
- Read each section in order on first pass; later sections depend on earlier transport and object-contract concepts.
- If you are primarily a researcher, start with the scientific workflow below and then return to the architecture sections only as needed.
- If you are building pipelines or package integrations, the later sections document the nextgen contracts in detail.
How to read each section:
- Each major section is written to answer four questions: what problem this part of the package solves, what object or table it returns, how to tell if the result is usable, and what the next step usually is.
- Each subsection follows the same pattern:
Purposetells you when to use a function, theMinimalexample shows the smallest valid call, theTypicalexample shows a realistic workflow call, theAdvancedexample shows how the function behaves in robust pipeline code, andInterpretationexplains how to read the returned object. - When an example returns a typed object rather than a plain table,
the first things to inspect are
success,error,pending,from_cache, and request metadata fromrequest_args(). - When an example returns a table, focus first on the row meaning, the
key identifier columns (
CID,SID,AID), and whether the output is already suitable for joining, plotting, or modeling.
Research-First Workflow
This section answers the practical question a typical PubChemR user starts with:
“Given a known compound, how do I triage related compounds by properties and assay activity?”
The example below uses an aspirin-like seed structure. In live mode it can query PubChem similarity results; in offline mode it falls back to exported deterministic example data so the workflow still runs end-to-end.
library(PubChemR)
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
library(tibble)
run_live <- identical(Sys.getenv("PUBCHEMR_RUN_LIVE"), "true")
`%||%` <- function(x, y) if (is.null(x)) y else x
safe_call <- function(expr) {
tryCatch(
expr,
error = function(e) structure(list(message = conditionMessage(e)), class = "pc_error")
)
}
summarize_any <- function(x) {
if (inherits(x, "pc_error")) {
return(tibble(ok = FALSE, class = "pc_error", note = x$message, rows = NA_integer_, cols = NA_integer_))
}
if (inherits(x, "PubChemResult")) {
return(tibble(
ok = isTRUE(x$success),
class = class(x)[1],
note = if (isTRUE(x$success)) "success" else (x$error$code %||% "error"),
rows = nrow(as_tibble(x)),
cols = ncol(as_tibble(x))
))
}
if (inherits(x, "PubChemAsyncQuery")) {
return(tibble(ok = TRUE, class = class(x)[1], note = "async query object", rows = NA_integer_, cols = NA_integer_))
}
if (is.data.frame(x)) {
return(tibble(ok = TRUE, class = class(x)[1], note = "tabular output", rows = nrow(x), cols = ncol(x)))
}
tibble(ok = TRUE, class = class(x)[1], note = "non-tabular output", rows = NA_integer_, cols = NA_integer_)
}Use Case
Goal:
- Start from a known analgesic scaffold.
- Resolve or reuse a small candidate set.
- Retrieve or reuse physicochemical properties.
- Attach assay activity evidence.
- Filter to compounds that look drug-like and show any evidence of activity.
research_seed_smiles <- "CC(=O)OC1=CC=CC=C1C(=O)O"
assay_payload <- pc_example_assaysummary_payload()
feature_tbl_synthetic <- pc_example_feature_table() %>%
mutate(
CanonicalSMILES = c(
"CC(=O)OC1=CC=CC=C1C(=O)O",
"CC(C)CC1=CC=C(C=C1)C(C)C(=O)O",
"CN1C=NC2=C1C(=O)N(C(=O)N2)C"
)
)
research_similarity <- if (run_live) {
pc_similarity_search(
identifier = research_seed_smiles,
namespace = "smiles",
threshold = 90,
max_records = 25,
cache = TRUE
)
} else {
pc_similarity_search(
identifier = research_seed_smiles,
namespace = "smiles",
threshold = 90,
max_records = 25,
cache = TRUE,
offline = TRUE
)
}
research_cids <- if (inherits(research_similarity, "PubChemResult") && isTRUE(research_similarity$success)) {
as_tibble(research_similarity) %>%
filter(!is.na(CID)) %>%
transmute(CID = as.character(CID)) %>%
distinct()
} else {
feature_tbl_synthetic %>%
transmute(CID)
}
research_features_try <- pc_feature_table(
identifier = research_cids$CID,
properties = c("MolecularWeight", "XLogP", "TPSA", "HBondDonorCount", "HBondAcceptorCount"),
namespace = "cid",
cache = TRUE,
offline = !run_live,
error_mode = "result"
)
research_features <- if (inherits(research_features_try, "PubChemResult")) {
feature_tbl_synthetic
} else {
research_features_try %>%
mutate(CID = as.character(CID))
}
research_assay_long <- pc_assay_activity_long(
x = assay_payload,
add_outcome_value = TRUE
)
research_activity_summary <- research_assay_long %>%
group_by(CID) %>%
summarise(
n_assays = n(),
outcomes = paste(sort(unique(ActivityOutcome)), collapse = ", "),
any_active = any(ActivityOutcomeValue == 1, na.rm = TRUE),
median_activity_uM = if (all(is.na(ActivityValue_uM))) NA_real_ else median(ActivityValue_uM, na.rm = TRUE),
.groups = "drop"
)
research_triage <- research_features %>%
left_join(research_activity_summary, by = "CID") %>%
mutate(
n_assays = ifelse(is.na(n_assays), 0L, as.integer(n_assays)),
any_active = ifelse(is.na(any_active), FALSE, any_active),
outcomes = ifelse(is.na(outcomes), "No assay evidence", outcomes),
passes_rule_of_five_like =
MolecularWeight <= 500 &
XLogP <= 5 &
HBondDonorCount <= 5 &
HBondAcceptorCount <= 10,
activity_bucket = case_when(
any_active ~ "Any Active",
n_assays > 0 ~ "Assayed, no active hit",
TRUE ~ "No assay evidence in fixture"
)
) %>%
arrange(desc(any_active), desc(passes_rule_of_five_like), MolecularWeight)
research_triage %>%
select(CID, MolecularWeight, XLogP, TPSA, n_assays, any_active, passes_rule_of_five_like, activity_bucket)
#> # A tibble: 3 × 8
#> CID MolecularWeight XLogP TPSA n_assays any_active passes_rule_of_five_like
#> <chr> <dbl> <dbl> <dbl> <int> <lgl> <lgl>
#> 1 2244 180. 1.2 63.6 2 TRUE TRUE
#> 2 5957 194. 0.1 86.6 0 FALSE TRUE
#> 3 3672 206. 3.5 37.3 1 FALSE TRUE
#> # ℹ 1 more variable: activity_bucket <chr>What this shows:
- PubChemR can support a familiar medicinal-chemistry screening pattern: seed compound -> candidate set -> properties -> assay evidence -> triage.
- The same code path can be live or deterministic depending on
run_live. - The nextgen layer is useful here because data acquisition can fail while the workflow still remains explicit and reproducible.
Filter and Rank
research_ranked <- research_triage %>%
filter(passes_rule_of_five_like) %>%
mutate(
potency_score = ifelse(is.na(median_activity_uM), 0, 1 / (1 + median_activity_uM)),
rank_score =
2 * as.numeric(any_active) +
potency_score +
0.10 * n_assays -
abs(XLogP - 2.5) / 5
) %>%
arrange(desc(rank_score), MolecularWeight) %>%
mutate(rank = row_number())
list(
total_candidates = nrow(research_triage),
druglike_candidates = nrow(research_ranked),
top_candidates = research_ranked %>%
select(rank, CID, MolecularWeight, XLogP, n_assays, any_active, median_activity_uM, rank_score)
)
#> $total_candidates
#> [1] 3
#>
#> $druglike_candidates
#> [1] 3
#>
#> $top_candidates
#> # A tibble: 3 × 8
#> rank CID MolecularWeight XLogP n_assays any_active median_activity_uM
#> <int> <chr> <dbl> <dbl> <int> <lgl> <dbl>
#> 1 1 2244 180. 1.2 2 TRUE 1.2
#> 2 2 3672 206. 3.5 1 FALSE 12.5
#> 3 3 5957 194. 0.1 0 FALSE NA
#> # ℹ 1 more variable: rank_score <dbl>Interpretation:
- The workflow now completes the usual screening loop: retrieve, filter, summarize activity, then rank candidates.
- The ranking formula is intentionally simple and transparent; it is a vignette-level triage score, not a production QSAR model.
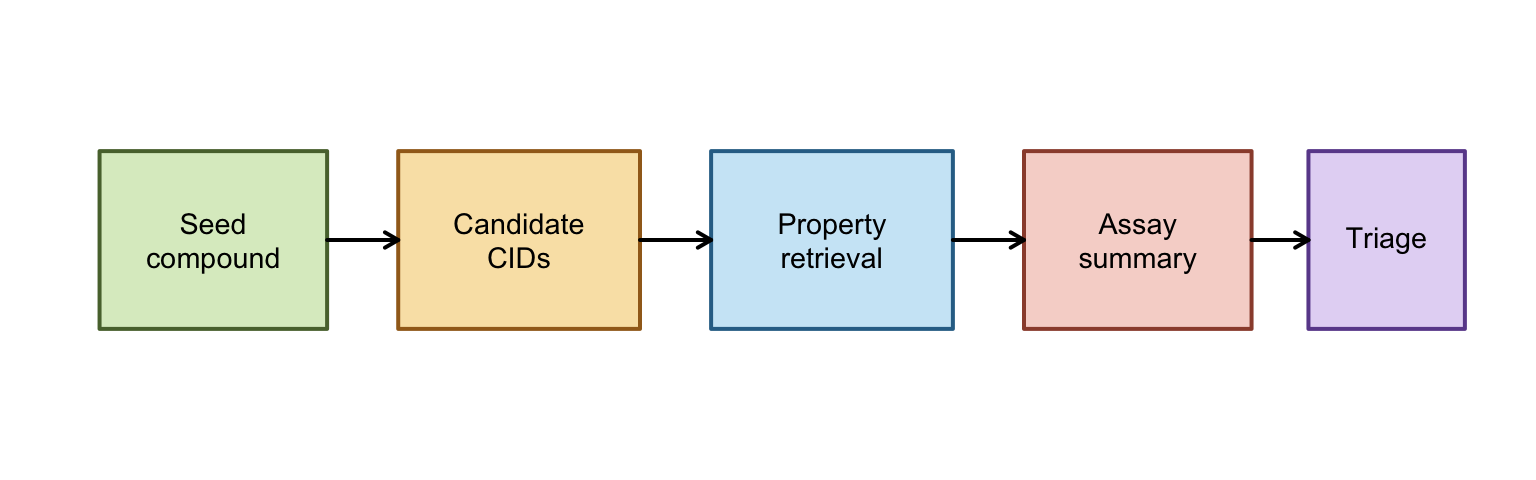
Pipeline Diagram
Interpretation:
- This diagram is the mental model for the rest of the vignette: each box is a stage where PubChemR changes the shape of the data, not just retrieves it.
- The early boxes are retrieval-focused, the middle boxes are normalization and summarization, and the final box is where scientific judgment enters through filtering and ranking.
- If a later section feels abstract, map it back to one box here and ask what that stage must return for the next stage to work.
Quick Plots
op <- par(mfrow = c(2, 2), mar = c(4, 4, 2, 1))
hist(
research_triage$MolecularWeight,
main = "Candidate Molecular Weight",
xlab = "MolecularWeight",
col = "#CDE8F6",
border = "white"
)
barplot(
table(research_assay_long$ActivityOutcome),
main = "Activity Outcome Counts",
ylab = "Count",
las = 2,
col = c("#DCECC9", "#F9E3B4", "#F6D6D0")
)
plot(
research_triage$XLogP,
research_triage$MolecularWeight,
pch = 19,
col = ifelse(research_triage$any_active, "#58713A", "#9B4D3A"),
xlab = "XLogP",
ylab = "MolecularWeight",
main = "Property Space by Activity"
)
text(
research_triage$XLogP,
research_triage$MolecularWeight,
labels = research_triage$CID,
pos = 3,
cex = 0.8
)
barplot(
research_ranked$rank_score,
names.arg = research_ranked$CID,
main = "Ranked Candidate Scores",
ylab = "rank_score",
col = "#E4D8F5",
las = 2
)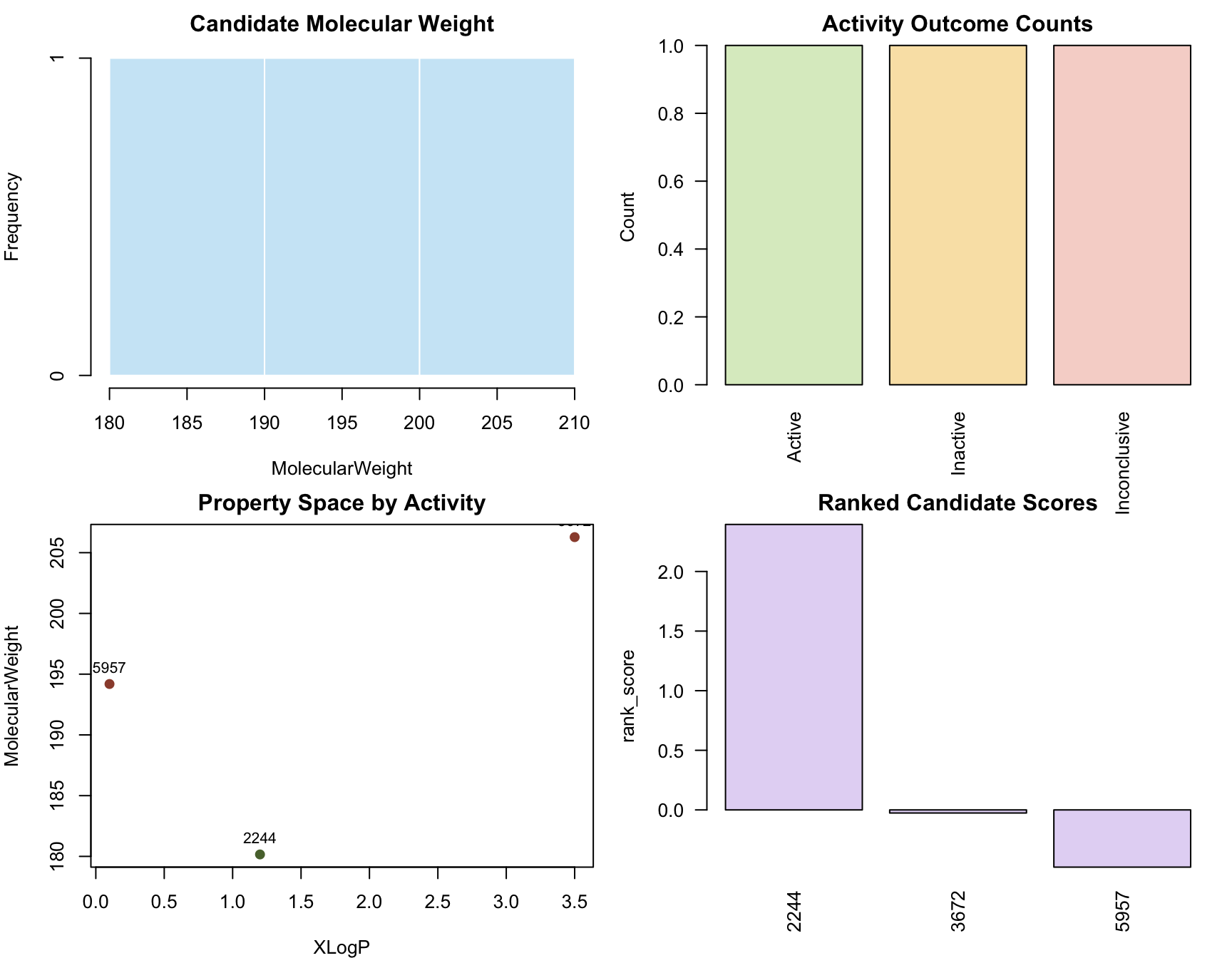
par(op)Interpretation:
- The histogram answers whether the candidate set sits in a plausible molecular weight range for small-molecule screening.
- The activity bar chart answers whether the assay evidence is informative or mostly empty/inconclusive.
- The property-space scatter plot answers whether active and inactive compounds occupy obviously different regions of simple descriptor space.
- The ranked-score plot answers whether the workflow produces a usable ordering rather than only a raw collection of records.
Why It Matters
The rest of this vignette explains the machinery behind the workflow above:
- how requests are represented as typed objects,
- how cache/offline policies are controlled,
- how async and batch workflows are recovered safely,
- and how the analysis helpers connect retrieval to model-ready data.
Roadmap
The next-generation API in PubChemR centers around typed result objects, policy-controlled transport, orchestration helpers, and analysis-layer tools.
Read this section as a map, not as something you need to memorize. Its purpose is to show where each function family belongs in a real workflow and why the same package contains both scientific helpers and infrastructure helpers.
nextgen_function_map <- tibble(
family = c(
rep("Transport", 6),
rep("Wrappers", 7),
rep("Async", 3),
rep("Batch/Benchmark", 4),
rep("Analysis", 10),
rep("Helpers", 6)
),
function_name = c(
"pc_profile", "pc_config", "pc_request", "pc_response", "pc_cache_info/pc_cache_clear", "pc_capabilities",
"pc_compound", "pc_substance", "pc_assay", "pc_property", "pc_identifier_map", "pc_similarity_search", "pc_sdq_bioactivity",
"pc_submit", "pc_poll", "pc_collect",
"pc_batch", "pc_resume_batch", "pc_benchmark", "pc_benchmark_harness",
"pc_assay_activity_long", "pc_activity_outcome_map", "pc_activity_matrix", "pc_cross_domain_join",
"pc_feature_table", "pc_model_matrix", "pc_export_model_data", "pc_to_rcdk", "pc_to_chemminer", "pc_lifecycle_policy",
"pc_example_assaysummary_payload", "pc_example_feature_table", "as_tibble", "request_args", "has_hits", "retrieve"
)
)
nextgen_function_map
#> # A tibble: 36 × 2
#> family function_name
#> <chr> <chr>
#> 1 Transport pc_profile
#> 2 Transport pc_config
#> 3 Transport pc_request
#> 4 Transport pc_response
#> 5 Transport pc_cache_info/pc_cache_clear
#> 6 Transport pc_capabilities
#> 7 Wrappers pc_compound
#> 8 Wrappers pc_substance
#> 9 Wrappers pc_assay
#> 10 Wrappers pc_property
#> # ℹ 26 more rowsWhat to expect:
- This table is the roadmap for the rest of the vignette.
- Every function listed above appears in runnable examples.
- Live-network examples are explicitly guarded by
run_live. - Exported deterministic fixtures are used wherever possible so examples stay stable offline.
How to interpret the roadmap:
Transportfunctions control how requests are made and how failures are represented.Wrapperstell PubChem what kind of record you want.AsyncandBatch/Benchmarkfunctions become relevant when a workflow must scale or recover from interruptions.Analysisfunctions turn retrieved data into tables, matrices, and exports a scientist can use directly.Helpersare the glue for inspecting, standardizing, and reusing outputs.
Core object classes used throughout:
PubChemResult: typed transport result (success,data,error, metadata).PubChemRecord/PubChemIdMap: specializedPubChemResultsubclasses.PubChemAsyncQuery: async container for submit/poll/collect workflows.PubChemBatchResult: chunk-level execution summary and checkpoint metadata.PubChemBenchmarkReport: scenario-level benchmark summary with pass/fail gates.PubChemModelMatrix/PubChemSparseActivityMatrix: modeling-ready matrix wrappers.
Setup
A robust nextgen workflow starts with explicit transport policy and deterministic runtime settings.
Section contract:
- Read this section before you trust any later example.
- Inputs: runtime environment, cache location, and transport defaults.
- Outputs: a known policy state and an auditable capability snapshot.
- Interpret the outputs by asking: where is the cache, is live mode allowed, and do optional packages exist in this session.
- Pitfalls: hidden global-state mutations and accidental live requests.
- Performance note: stable cache policy is usually more important than raw request speed in iterative workflows.
cfg_before <- pc_config()
cfg_before
#> $rate_limit
#> [1] 5
#>
#> $timeout
#> [1] 60
#>
#> $retries
#> [1] 3
#>
#> $pause_base
#> [1] 1
#>
#> $pause_cap
#> [1] 8
#>
#> $user_agent
#> [1] "PubChemR/3.0.0"
#>
#> $cache_dir
#> [1] "/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T//RtmpHAl5DI/PubChemR_cache"
#>
#> $cache_ttl
#> [1] 86400
#>
#> $offline
#> [1] FALSE
pc_profile("default")
#> $rate_limit
#> [1] 5
#>
#> $timeout
#> [1] 60
#>
#> $retries
#> [1] 3
#>
#> $pause_base
#> [1] 1
#>
#> $pause_cap
#> [1] 8
#>
#> $user_agent
#> [1] "PubChemR/3.0.0"
#>
#> $cache_dir
#> [1] "/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T//RtmpHAl5DI/PubChemR_cache"
#>
#> $cache_ttl
#> [1] 86400
#>
#> $offline
#> [1] FALSE
cfg_after_profile <- pc_config()
cfg_after_profile[c("rate_limit", "timeout", "retries", "cache_ttl")]
#> $rate_limit
#> [1] 5
#>
#> $timeout
#> [1] 60
#>
#> $retries
#> [1] 3
#>
#> $cache_ttl
#> [1] 86400
# Keep cache in a temp path during vignette execution.
work_cache <- file.path(tempdir(), "pubchemr-vignette-cache")
dir.create(work_cache, recursive = TRUE, showWarnings = FALSE)
pc_config(cache_dir = work_cache, offline = FALSE)
#> $rate_limit
#> [1] 5
#>
#> $timeout
#> [1] 60
#>
#> $retries
#> [1] 3
#>
#> $pause_base
#> [1] 1
#>
#> $pause_cap
#> [1] 8
#>
#> $user_agent
#> [1] "PubChemR/3.0.0"
#>
#> $cache_dir
#> [1] "/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T//RtmpjLjM6G/pubchemr-vignette-cache"
#>
#> $cache_ttl
#> [1] 86400
#>
#> $offline
#> [1] FALSE
pc_cache_clear(cache_dir = work_cache, memory = TRUE, disk = TRUE)
pc_cache_info(cache_dir = work_cache)
#> # A tibble: 1 × 4
#> memory_entries disk_entries disk_size_bytes cache_dir
#> <int> <int> <dbl> <chr>
#> 1 0 0 0 /var/folders/dj/y28dp44x303ggfg6r…What to expect:
pc_profile()applies a baseline policy.pc_config()confirms active runtime state.- Cache is isolated to
tempdir()to avoid modifying user directories.
pc_capabilities(cache_dir = work_cache, check_network = FALSE)
#> # A tibble: 1 × 9
#> live_enabled offline_mode cache_dir cache_exists cache_writable optional_rcdk
#> <lgl> <lgl> <chr> <lgl> <lgl> <lgl>
#> 1 TRUE FALSE /var/fold… TRUE TRUE FALSE
#> # ℹ 3 more variables: optional_chemminer <lgl>, optional_matrix <lgl>,
#> # network_reachable <lgl>Interpretation:
pc_capabilities()gives a preflight summary for optional dependencies, cache writability, and offline/live mode.- Use this as an early diagnostic before long workflows or CI jobs.
Key pitfalls and notes:
rate_limitandcache_ttlare validated; invalid values fail early.- Use explicit cache directories in scripts for deterministic CI behavior.
- Restore old config in project code when changing globals
(
pc_config(...)).
Core Contracts
The transport contract is the foundation for every other nextgen function.
If results feel abstract later in the vignette, come back here. This section defines what a “good” result looks like, what a controlled failure looks like, and why the package often returns objects rather than immediately returning a data frame.
Parsing Results
Purpose:
- Normalize success/fault payloads into a stable structure
(
success,data,error, metadata).
Minimal example
ok_text <- '{"PropertyTable":{"Properties":[{"CID":2244,"MolecularWeight":180.16}]}}'
res_ok <- pc_response(
ok_text,
request = list(domain = "compound", namespace = "cid", identifier = 2244, operation = "property/MolecularWeight")
)
class(res_ok)
#> [1] "PubChemResult"
res_ok$success
#> [1] TRUE
as_tibble(res_ok)
#> # A tibble: 1 × 6
#> success status from_cache pending CID MolecularWeight
#> <lgl> <int> <lgl> <lgl> <dbl> <dbl>
#> 1 TRUE NA FALSE FALSE 2244 180.Interpretation:
successisTRUEandas_tibble()exposes payload content as a flat table.
Typical example
fault_text <- '{"Fault":{"Code":"PUGREST.BadRequest","Message":"Invalid request"}}'
res_fault <- pc_response(fault_text, request = list(domain = "compound"))
res_fault$success
#> [1] FALSE
res_fault$error$code
#> [1] "PUGREST.BadRequest"
res_fault$error$message
#> [1] "PUGREST.BadRequest" "Invalid request"Interpretation:
- Fault payloads remain structured; they do not crash the pipeline by default.
- Downstream code can branch on
successanderror$code.
Advanced example
waiting_text <- '{"Waiting":{"ListKey":"example-listkey"}}'
res_wait <- pc_response(waiting_text, request = list(domain = "compound", operation = "cids"))
tibble(
success = res_wait$success,
pending = res_wait$pending,
listkey = res_wait$listkey
)
#> # A tibble: 1 × 3
#> success pending listkey
#> <lgl> <lgl> <chr>
#> 1 TRUE TRUE example-listkeyInterpretation:
- Pending responses carry
listkey, enabling async flow (pc_poll()/pc_collect()).
Performance and reproducibility notes:
pc_response()is local parsing, so it is deterministic and fast.- It is ideal for unit-testing workflow logic without network dependency.
Inspecting Results
Purpose:
- Inspect request provenance (
request_args()), convert typed results (as_tibble()), and detect hit availability (has_hits()).
Minimal example
request_args(res_ok)
#> $domain
#> [1] "compound"
#>
#> $namespace
#> [1] "cid"
#>
#> $identifier
#> [1] 2244
#>
#> $operation
#> [1] "property/MolecularWeight"
request_args(res_ok, "identifier")
#> [1] 2244Interpretation:
- Request metadata stays attached to result objects for provenance auditing.
Typical example
id_text <- '{"IdentifierList":{"CID":[2244,3672,5957]}}'
id_res <- pc_response(id_text, request = list(domain = "compound", namespace = "name", identifier = "aspirin", operation = "cids"))
as_tibble(id_res)
#> # A tibble: 3 × 5
#> success status from_cache pending CID
#> <lgl> <int> <lgl> <lgl> <dbl>
#> 1 TRUE NA FALSE FALSE 2244
#> 2 TRUE NA FALSE FALSE 3672
#> 3 TRUE NA FALSE FALSE 5957Interpretation:
as_tibble()handles common payload families (PropertyTable,IdentifierList,InformationList).
Advanced example
if (exists("has_hits", mode = "function")) {
synthetic_hit_flags <- structure(
list(
request_args = list(identifier = c("aspirin", "unknown")),
has_hits = c(TRUE, FALSE),
success = c(TRUE, FALSE)
),
class = "PubChemInstance_CIDs"
)
has_hits(synthetic_hit_flags)
} else {
"has_hits() is unavailable in this installed package build."
}
#> aspirin unknown
#> TRUE FALSEInterpretation:
has_hits()is useful when output formatting depends on non-empty match sets.
Legacy Access
Purpose:
- Provide a legacy-style accessor workflow directly on nextgen typed results.
- Extract either the full payload table (
.slot = "data") or nested fields (.slot = "IdentifierList/CID",.slot = "error/code").
Minimal example
retrieve(id_res)
#> # A tibble: 3 × 1
#> CID
#> <dbl>
#> 1 2244
#> 2 3672
#> 3 5957Interpretation:
- For successful
PubChemResult, default retrieval returns payload data in a tabular form when possible.
Typical example
retrieve(id_res, .slot = "IdentifierList/CID", .to.data.frame = FALSE)
#> [1] 2244 3672 5957Interpretation:
- Nested extraction supports slash-delimited paths for concise access to deeply nested payload fields.
Advanced example
retrieve(res_fault, .slot = "error/code", .to.data.frame = FALSE)
#> [1] "PUGREST.BadRequest"Interpretation:
- Retrieval from error paths is useful when building deterministic failure routing or metrics around specific error codes.
Object Surfaces
Purpose:
- Inspect the stable object surfaces exposed by the nextgen API.
- Confirm that classes, print methods, and metadata fields are designed for programmatic branching as well as interactive inspection.
Minimal example
list(
result_class = class(res_ok),
result_request_fields = names(request_args(res_ok)),
result_error_fields = names(res_fault$error)
)
#> $result_class
#> [1] "PubChemResult"
#>
#> $result_request_fields
#> [1] "domain" "namespace" "identifier" "operation"
#>
#> $result_error_fields
#> [1] "code" "message" "status" "details"Typical example
capture.output(print(res_fault))
#> [1] ""
#> [2] " PubChemResult "
#> [3] ""
#> [4] " - Success: FALSE"
#> [5] " - Status: NA"
#> [6] " - Pending: FALSE"
#> [7] " - From cache: FALSE"
#> [8] " - Error Code: PUGREST.BadRequest"
#> [9] " - Error Message: PUGREST.BadRequestInvalid request"Advanced example
typed_surface <- list(
ok = res_ok[c("success", "status", "pending", "from_cache")],
fault = res_fault[c("success", "status", "pending")],
wait = res_wait[c("success", "pending", "listkey")]
)
typed_surface
#> $ok
#> $ok$success
#> [1] TRUE
#>
#> $ok$status
#> [1] NA
#>
#> $ok$pending
#> [1] FALSE
#>
#> $ok$from_cache
#> [1] FALSE
#>
#>
#> $fault
#> $fault$success
#> [1] FALSE
#>
#> $fault$status
#> [1] NA
#>
#> $fault$pending
#> [1] FALSE
#>
#>
#> $wait
#> $wait$success
#> [1] TRUE
#>
#> $wait$pending
#> [1] TRUE
#>
#> $wait$listkey
#> [1] "example-listkey"Interpretation:
- Print methods are concise by design; use fields like
success,pending,listkey,status, anderrorfor automation. request_args()is the stable provenance interface when objects move between orchestration, analysis, and export steps.
Transport
This section covers pc_profile(),
pc_config(), pc_capabilities(),
pc_request(), pc_cache_info(), and
pc_cache_clear() with escalating complexity.
Section contract:
- Use this section to understand how PubChemR talks to PubChem, not to analyze chemistry yet.
- Inputs: profile names, transport overrides, request descriptors, and cache policy.
- Outputs: typed
PubChemResultobjects and cache diagnostics. - Interpret the outputs by checking whether the request succeeded, whether the payload came from cache, and whether the failure is a transport problem or a domain problem.
- Pitfalls: mixing offline mode with uncached requests; overly
aggressive
force_refreshin reproducible workflows. - Performance note: tune
rate_limit,timeout, and cache policy together; tuning only one parameter rarely yields stable gains.
Profiles
Purpose:
- Apply a named policy template (
default,cloud,high_throughput). - Typical output: a named list matching active transport config fields.
Minimal example
pc_profile("default")$rate_limit
#> [1] 5Typical example
pc_profile("cloud", retries = 4, pause_cap = 12)[c("rate_limit", "retries", "pause_cap")]
#> $rate_limit
#> [1] 3
#>
#> $retries
#> [1] 4
#>
#> $pause_cap
#> [1] 12Advanced example
old_cfg <- pc_config()
pc_profile("high_throughput", cache_ttl = 3600, timeout = 45)
#> $rate_limit
#> [1] 10
#>
#> $timeout
#> [1] 45
#>
#> $retries
#> [1] 4
#>
#> $pause_base
#> [1] 0.5
#>
#> $pause_cap
#> [1] 10
#>
#> $user_agent
#> [1] "PubChemR/3.0.0"
#>
#> $cache_dir
#> [1] "/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T//RtmpjLjM6G/pubchemr-vignette-cache"
#>
#> $cache_ttl
#> [1] 3600
#>
#> $offline
#> [1] FALSE
new_cfg <- pc_config()
list(
before = old_cfg[c("rate_limit", "timeout", "cache_ttl")],
after = new_cfg[c("rate_limit", "timeout", "cache_ttl")]
)
#> $before
#> $before$rate_limit
#> [1] 3
#>
#> $before$timeout
#> [1] 120
#>
#> $before$cache_ttl
#> [1] 604800
#>
#>
#> $after
#> $after$rate_limit
#> [1] 10
#>
#> $after$timeout
#> [1] 45
#>
#> $after$cache_ttl
#> [1] 3600Interpretation:
- Profiles are starting points; explicit overrides should be documented in production code.
- For reproducibility, capture profile + overrides together in project config.
Configuration
Purpose:
- Read or update global transport defaults.
- Typical output: a complete config list including
offline, retries, and cache policy.
Minimal example
pc_config()[c("rate_limit", "timeout", "cache_ttl", "offline")]
#> $rate_limit
#> [1] 10
#>
#> $timeout
#> [1] 45
#>
#> $cache_ttl
#> [1] 3600
#>
#> $offline
#> [1] FALSETypical example
pc_config(rate_limit = 6, timeout = 50, cache_ttl = 1800, offline = FALSE)
#> $rate_limit
#> [1] 6
#>
#> $timeout
#> [1] 50
#>
#> $retries
#> [1] 4
#>
#> $pause_base
#> [1] 0.5
#>
#> $pause_cap
#> [1] 10
#>
#> $user_agent
#> [1] "PubChemR/3.0.0"
#>
#> $cache_dir
#> [1] "/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T//RtmpjLjM6G/pubchemr-vignette-cache"
#>
#> $cache_ttl
#> [1] 1800
#>
#> $offline
#> [1] FALSE
pc_config()[c("rate_limit", "timeout", "cache_ttl", "offline")]
#> $rate_limit
#> [1] 6
#>
#> $timeout
#> [1] 50
#>
#> $cache_ttl
#> [1] 1800
#>
#> $offline
#> [1] FALSEAdvanced example
safe_call(pc_config(rate_limit = 0))
#> $message
#> [1] "'rate_limit' must be a finite numeric scalar > 0."
#>
#> attr(,"class")
#> [1] "pc_error"Interpretation:
- Invalid config values are rejected immediately, reducing hidden runtime failures.
pc_config()mutates package-level state, so scripts should set and restore it intentionally.
Capabilities
Purpose:
- Preflight the runtime before doing transport, caching, optional bridge, or live-network work.
- Typical output: one-row tibble containing cache, dependency, and connectivity flags.
Minimal example
pc_capabilities(cache_dir = work_cache, check_network = FALSE)
#> # A tibble: 1 × 9
#> live_enabled offline_mode cache_dir cache_exists cache_writable optional_rcdk
#> <lgl> <lgl> <chr> <lgl> <lgl> <lgl>
#> 1 TRUE FALSE /var/fold… TRUE TRUE FALSE
#> # ℹ 3 more variables: optional_chemminer <lgl>, optional_matrix <lgl>,
#> # network_reachable <lgl>Typical example
pc_capabilities(
cache_dir = work_cache,
check_network = run_live,
network_timeout = 2
)
#> # A tibble: 1 × 9
#> live_enabled offline_mode cache_dir cache_exists cache_writable optional_rcdk
#> <lgl> <lgl> <chr> <lgl> <lgl> <lgl>
#> 1 TRUE FALSE /var/fold… TRUE TRUE FALSE
#> # ℹ 3 more variables: optional_chemminer <lgl>, optional_matrix <lgl>,
#> # network_reachable <lgl>Interpretation:
check_network = FALSEkeeps examples deterministic.- In interactive sessions,
check_network = TRUEis a practical early warning before running longer live workflows.
Requests
Purpose:
- Unified low-level transport for all
pc_*wrappers. - Typical output:
PubChemResultwithsuccess,error, and replay metadata.
Minimal example
req_min <- pc_request(identifier = 2244, offline = TRUE, cache = TRUE)
summarize_any(req_min)
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemResult OfflineCacheMiss 1 4Interpretation:
- In a clean offline session, this should return a structured
OfflineCacheMissfailure.
Typical example
req_typed <- pc_request(
domain = "compound",
namespace = "cid",
identifier = 2244,
operation = "property/MolecularWeight,XLogP",
output = "JSON",
cache = TRUE,
offline = TRUE
)
summarize_any(req_typed)
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemResult OfflineCacheMiss 1 4Interpretation:
- Even on failure, metadata remains complete for diagnostics.
Advanced example
req_post <- pc_request(
domain = "compound",
namespace = "smiles",
identifier = "CCO",
operation = "cids",
method = "POST",
body = list(smiles = "CCO"),
output = "JSON",
cache = TRUE,
offline = TRUE
)
list(
summary = summarize_any(req_post),
request = request_args(req_post)[c("method", "namespace", "identifier", "operation")]
)
#> $summary
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemResult OfflineCacheMiss 1 4
#>
#> $request
#> $request$method
#> [1] "POST"
#>
#> $request$namespace
#> [1] "smiles"
#>
#> $request$identifier
#> [1] "CCO"
#>
#> $request$operation
#> [1] "cids"Interpretation:
pc_request()supports bothGETandPOST; wrappers inherit that transport surface through....- For deterministic tests, prefer
offline = TRUEand assert on structured error codes.
Cache State
Purpose:
- Observe and reset cache state.
- Typical output: one-row tibble diagnostics from
pc_cache_info().
Minimal example
pc_cache_info(cache_dir = work_cache)
#> # A tibble: 1 × 4
#> memory_entries disk_entries disk_size_bytes cache_dir
#> <int> <int> <dbl> <chr>
#> 1 0 0 0 /var/folders/dj/y28dp44x303ggfg6r…Typical example
pc_cache_clear(cache_dir = work_cache, memory = TRUE, disk = FALSE)
pc_cache_info(cache_dir = work_cache)
#> # A tibble: 1 × 4
#> memory_entries disk_entries disk_size_bytes cache_dir
#> <int> <int> <dbl> <chr>
#> 1 0 0 0 /var/folders/dj/y28dp44x303ggfg6r…Advanced example
pc_cache_clear(cache_dir = work_cache, memory = TRUE, disk = TRUE)
pc_cache_info(cache_dir = work_cache)
#> # A tibble: 1 × 4
#> memory_entries disk_entries disk_size_bytes cache_dir
#> <int> <int> <dbl> <chr>
#> 1 0 0 0 /var/folders/dj/y28dp44x303ggfg6r…Interpretation:
- Use full cache clears before controlled benchmarks.
- Avoid unnecessary clears in production runs where caching is intentional.
- Pair cache diagnostics with
pc_capabilities()when debugging environment-specific behavior.
Domain Wrappers
These wrappers express intent while preserving the
PubChemResult contract. pc_similarity_search()
and pc_sdq_bioactivity() are introduced here, then
revisited in more detail in the analysis section because they often feed
downstream modeling workflows.
Section contract:
- Use this section when you know what you want from PubChem conceptually (compound, assay, property, mapping) and need to know which wrapper to call.
- Inputs: domain-specific identifiers, namespaces, operations/search types, and transport controls.
- Outputs: typed wrapper results
(
PubChemRecord/PubChemIdMap) or table-shaped outputs for SDQ and higher-level analysis wrappers. - Interpret the outputs first by object class, then by payload family: record-style results keep structure, ID-mapping results usually become small tables, and SDQ results can already be analysis-ready tibbles.
- Pitfalls: namespace/identifier mismatches and expecting identical payload schema across wrapper families.
- Performance note: use wrapper-level cache settings for repeated reads in exploration; move to staged persisted files for modeling.
Minimal Calls
wrapper_minimal <- list(
pc_compound = safe_call(pc_compound(2244, offline = TRUE)),
pc_substance = safe_call(pc_substance(5360534, offline = TRUE)),
pc_assay = safe_call(pc_assay(367, offline = TRUE)),
pc_property = safe_call(pc_property(2244, properties = "MolecularWeight", offline = TRUE)),
pc_identifier_map = safe_call(pc_identifier_map("aspirin", namespace = "name", to = "cids", offline = TRUE)),
pc_similarity_search = safe_call(pc_similarity_search("CCO", namespace = "smiles", offline = TRUE)),
pc_sdq_bioactivity = if (run_live) {
safe_call(pc_sdq_bioactivity(2244, rate_limit = FALSE, limit = 200L))
} else {
structure(list(message = "Skipped (set PUBCHEMR_RUN_LIVE=true to run SDQ live examples)."), class = "pc_error")
}
)
bind_rows(lapply(wrapper_minimal, summarize_any), .id = "function")
#> # A tibble: 7 × 6
#> `function` ok class note rows cols
#> <chr> <lgl> <chr> <chr> <int> <int>
#> 1 pc_compound FALSE PubChemRecord OfflineCacheMi… 1 4
#> 2 pc_substance FALSE PubChemRecord OfflineCacheMi… 1 4
#> 3 pc_assay FALSE PubChemRecord OfflineCacheMi… 1 4
#> 4 pc_property FALSE PubChemRecord OfflineCacheMi… 1 4
#> 5 pc_identifier_map FALSE PubChemIdMap OfflineCacheMi… 1 5
#> 6 pc_similarity_search FALSE PubChemSimilarityResult OfflineCacheMi… 1 5
#> 7 pc_sdq_bioactivity FALSE pc_error Skipped (set P… NA NAInterpretation:
- In offline mode, wrappers return typed failures rather than hard crashes.
pc_sdq_bioactivity()is inherently live-network; this vignette guards it.- Use
retrieve()oras_tibble()immediately after wrapper calls to normalize downstream expectations.
Typical Calls
wrapper_typical <- list(
pc_compound = safe_call(pc_compound(identifier = c(2244, 3672), namespace = "cid", operation = "record", offline = TRUE)),
pc_substance = safe_call(pc_substance(identifier = c(5360534, 5360535), namespace = "sid", operation = "record", offline = TRUE)),
pc_assay = safe_call(pc_assay(identifier = c(367, 2551), namespace = "aid", operation = "description", offline = TRUE)),
pc_property = safe_call(pc_property(identifier = c(2244, 3672), properties = c("MolecularWeight", "XLogP"), namespace = "cid", offline = TRUE)),
pc_identifier_map = safe_call(pc_identifier_map(identifier = c("aspirin", "caffeine"), namespace = "name", to = "cids", domain = "compound", offline = TRUE)),
pc_similarity_search = safe_call(pc_similarity_search(identifier = "CC(=O)OC1=CC=CC=C1C(=O)O", namespace = "smiles", threshold = 90, max_records = 25, offline = TRUE)),
pc_sdq_bioactivity = if (run_live) {
safe_call(pc_sdq_bioactivity(identifier = 2244, namespace = "cid", limit = 200L, order = "activity,asc", rate_limit = FALSE))
} else {
structure(list(message = "Skipped (no live network)."), class = "pc_error")
}
)
bind_rows(lapply(wrapper_typical, summarize_any), .id = "function")
#> # A tibble: 7 × 6
#> `function` ok class note rows cols
#> <chr> <lgl> <chr> <chr> <int> <int>
#> 1 pc_compound FALSE PubChemRecord OfflineCacheMi… 1 4
#> 2 pc_substance FALSE PubChemRecord OfflineCacheMi… 1 4
#> 3 pc_assay FALSE PubChemRecord OfflineCacheMi… 1 4
#> 4 pc_property FALSE PubChemRecord OfflineCacheMi… 1 4
#> 5 pc_identifier_map FALSE PubChemIdMap OfflineCacheMi… 1 5
#> 6 pc_similarity_search FALSE PubChemSimilarityResult OfflineCacheMi… 1 5
#> 7 pc_sdq_bioactivity FALSE pc_error Skipped (no li… NA NAInterpretation:
- Typical examples use realistic argument sets and explicit namespaces.
- For live wrappers, keep limits small in interactive use and CI.
- Prefer explicit
operationandtoarguments over defaults in shared code.
Advanced Calls
wrapper_advanced <- list(
pc_compound = safe_call(pc_compound(identifier = 2244, namespace = "cid", operation = "record", method = "GET", cache = TRUE, offline = TRUE)),
pc_substance = safe_call(pc_substance(identifier = 5360534, namespace = "sid", operation = "record", cache = TRUE, offline = TRUE)),
pc_assay = safe_call(pc_assay(identifier = 367, namespace = "aid", operation = "summary", cache = TRUE, offline = TRUE)),
pc_property = safe_call(pc_property(identifier = c(2244, 3672), properties = c("MolecularWeight", "TPSA", "HBondDonorCount"), namespace = "cid", cache = TRUE, offline = TRUE)),
pc_identifier_map = safe_call(pc_identifier_map(identifier = c("aspirin", "ibuprofen", "caffeine"), namespace = "name", to = "cids", cache = TRUE, offline = TRUE)),
pc_similarity_search = safe_call(pc_similarity_search(identifier = "CCO", namespace = "smiles", searchtype = "fastsimilarity_2d", threshold = 85, max_records = 50, cache = TRUE, offline = TRUE)),
pc_sdq_bioactivity = if (run_live) {
safe_call(pc_sdq_bioactivity(identifier = 2244, namespace = "cid", collection = "bioactivity", limit = 500L, cache = TRUE, cache_dir = work_cache, cache_ttl = 3600, force_refresh = TRUE, rate_limit = FALSE))
} else {
structure(list(message = "Skipped (no live network)."), class = "pc_error")
}
)
bind_rows(lapply(wrapper_advanced, summarize_any), .id = "function")
#> # A tibble: 7 × 6
#> `function` ok class note rows cols
#> <chr> <lgl> <chr> <chr> <int> <int>
#> 1 pc_compound FALSE PubChemRecord OfflineCacheMi… 1 4
#> 2 pc_substance FALSE PubChemRecord OfflineCacheMi… 1 4
#> 3 pc_assay FALSE PubChemRecord OfflineCacheMi… 1 4
#> 4 pc_property FALSE PubChemRecord OfflineCacheMi… 1 4
#> 5 pc_identifier_map FALSE PubChemIdMap OfflineCacheMi… 1 5
#> 6 pc_similarity_search FALSE PubChemSimilarityResult OfflineCacheMi… 1 5
#> 7 pc_sdq_bioactivity FALSE pc_error Skipped (no li… NA NAInterpretation:
- Advanced usage combines explicit cache policy, parameter tuning, and reproducibility controls.
- Use
offline = TRUEfor deterministic local replay checks. pc_sdq_bioactivity()now supportserror_mode = "result"for typed failure handling.
Common wrapper pitfalls:
pc_property()requires at least one property name.pc_identifier_map()target must match the supported set (cids,sids,aids).pc_similarity_search()validates threshold and search mode.pc_sdq_bioactivity()is live-network by design; guard in reproducible docs and CI.
Async Queries
Purpose:
- Manage requests that may return listkeys and require polling.
Section contract:
- Inputs: an async-compatible request shape and polling policy
(
interval,max_attempts). - Outputs:
PubChemAsyncQueryobjects followed by terminalPubChemResultobjects. - Interpret the outputs by treating submit/poll/collect as separate states: submitted, still pending, or terminal.
- Pitfalls: polling indefinitely or ignoring
pending/listkeystatus. - Performance note: shorter intervals increase responsiveness but also request volume; use bounded attempts in CI.
Submit
q_min <- pc_submit(
domain = "compound",
namespace = "name",
identifier = "aspirin",
operation = "cids",
searchtype = "similarity",
options = list(Threshold = 90),
offline = TRUE
)
class(q_min)
#> [1] "PubChemAsyncQuery"
summarize_any(q_min$initial)
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemResult OfflineCacheMiss 1 4Interpretation:
pc_submit()always returns aPubChemAsyncQuerycontainer.
Poll
poll_min <- pc_poll("dummy-listkey", max_attempts = 1, interval = 0, offline = TRUE)
summarize_any(poll_min)
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemResult OfflineCacheMiss 1 4Interpretation:
- Polling returns typed failures when replay is unavailable.
- This is expected in a clean offline session.
Collect
collect_min <- pc_collect(q_min)
summarize_any(collect_min)
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemResult OfflineCacheMiss 1 4Interpretation:
pc_collect()centralizes branching logic (listkey vs no listkey).- For downstream pipelines, immediately convert terminal results with
retrieve(..., .slot = "data")oras_tibble().
Performance and reproducibility notes:
- Tune
intervalandmax_attemptsaccording to endpoint latency. - In CI, keep attempts small and assert error codes explicitly.
Batching and Benchmarks
This family includes pc_batch(),
pc_resume_batch(), pc_benchmark(), and
pc_benchmark_harness().
Section contract:
- Use this section when the question shifts from “can I retrieve one result?” to “can I run and recover this workflow at scale?”.
- Inputs: identifier vectors, deterministic worker functions, chunking policy, and optional checkpoint/report paths.
- Outputs: chunk-level execution objects and benchmark/harness summary tables.
- Interpret the outputs by focusing on chunk success ratios, resumability, and threshold pass/fail states rather than on the worker result values alone.
- Pitfalls: non-deterministic workers during checkpoint resume validation.
- Performance note: benchmark worker logic first with local deterministic functions, then swap in live transport.
Chunked Execution
Purpose:
- Chunk IDs, run a worker function, and preserve chunk-level success/error metadata.
Minimal example
batch_min <- pc_batch(
ids = 1:6,
fn = function(chunk_ids, ...) tibble(id = chunk_ids, score = chunk_ids * 10),
chunk_size = 2
)
batch_min
#>
#> PubChemBatchResult
#>
#> - Chunks: 3
#> - Chunk size: 2
#> - Parallel: FALSE
#> - Successful chunks: 3/3
as_tibble(batch_min)
#> # A tibble: 3 × 4
#> chunk n_ids success error
#> <int> <int> <lgl> <chr>
#> 1 1 2 TRUE ""
#> 2 2 2 TRUE ""
#> 3 3 2 TRUE ""Interpretation:
- Result includes
chunks, per-chunk success flags, and worker outputs.
Typical example (checkpointing)
cp_dir <- file.path(tempdir(), "pc-batch-checkpoints")
cp_id <- "vignette-batch-demo"
batch_cp <- pc_batch(
ids = 1:8,
fn = function(chunk_ids, ...) sum(chunk_ids),
chunk_size = 3,
checkpoint_dir = cp_dir,
checkpoint_id = cp_id
)
batch_cp$checkpoint
#> $enabled
#> [1] TRUE
#>
#> $id
#> [1] "vignette-batch-demo"
#>
#> $dir
#> [1] "/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T//RtmpjLjM6G/pc-batch-checkpoints"
#>
#> $manifest
#> [1] "/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T//RtmpjLjM6G/pc-batch-checkpoints/pc_batch_vignette-batch-demo_manifest.rds"
#>
#> $resumed
#> [1] FALSE
#>
#> $rerun_failed
#> [1] TRUEInterpretation:
- Checkpoint metadata tracks resumability and run identity.
Advanced example (parallel toggle + deterministic worker)
batch_adv <- pc_batch(
ids = letters[1:12],
fn = function(chunk_ids, ...) {
tibble(id = chunk_ids, nchar = nchar(chunk_ids), hash = as.integer(factor(chunk_ids)))
},
chunk_size = 4,
parallel = FALSE,
workers = 1
)
as_tibble(batch_adv)
#> # A tibble: 3 × 4
#> chunk n_ids success error
#> <int> <int> <lgl> <chr>
#> 1 1 4 TRUE ""
#> 2 2 4 TRUE ""
#> 3 3 4 TRUE ""Interpretation:
- Keep workers deterministic unless profiling throughput trade-offs.
Resume Runs
Purpose:
- Resume pending/failed chunk execution using checkpoint manifests.
Minimal example
resume_min <- pc_resume_batch(
fn = function(chunk_ids, ...) sum(chunk_ids),
checkpoint_dir = cp_dir,
checkpoint_id = cp_id
)
resume_min$checkpoint$resumed
#> [1] TRUETypical example
resume_typical <- pc_resume_batch(
fn = function(chunk_ids, ...) sum(chunk_ids),
checkpoint_dir = cp_dir,
checkpoint_id = cp_id,
rerun_failed = TRUE
)
as_tibble(resume_typical)
#> # A tibble: 3 × 4
#> chunk n_ids success error
#> <int> <int> <lgl> <chr>
#> 1 1 3 TRUE ""
#> 2 2 3 TRUE ""
#> 3 3 2 TRUE ""Advanced example
resume_adv <- pc_resume_batch(
fn = function(chunk_ids, ...) tibble(id = chunk_ids, value = as.numeric(chunk_ids)^2),
checkpoint_dir = cp_dir,
checkpoint_id = cp_id,
parallel = FALSE,
workers = 1,
rerun_failed = FALSE
)
resume_adv$chunk_status
#> [1] "success" "success" "success"Interpretation:
rerun_failedcontrols whether historical failed chunks are retried.
Benchmarking
Purpose:
- Compare chunking and parallel settings on elapsed-time and failure metrics.
Minimal example
bm_min <- pc_benchmark(
ids = 1:20,
fn = function(chunk_ids, ...) sum(chunk_ids),
chunk_sizes = c(5, 10),
parallel_options = FALSE
)
bm_min
#> # A tibble: 2 × 7
#> chunk_size parallel workers elapsed_sec chunks successful_chunks failed_chunks
#> <int> <lgl> <dbl> <dbl> <int> <int> <int>
#> 1 5 FALSE 1 0 4 4 0
#> 2 10 FALSE 1 0.00100 2 2 0Typical example
bm_typical <- pc_benchmark(
ids = 1:40,
fn = function(chunk_ids, ...) {
out <- sum(chunk_ids)
out
},
chunk_sizes = c(4, 8, 20),
parallel_options = c(FALSE)
)
bm_typical %>% arrange(elapsed_sec)
#> # A tibble: 3 × 7
#> chunk_size parallel workers elapsed_sec chunks successful_chunks failed_chunks
#> <int> <lgl> <dbl> <dbl> <int> <int> <int>
#> 1 20 FALSE 1 0 2 2 0
#> 2 4 FALSE 1 0.001000 10 10 0
#> 3 8 FALSE 1 0.00100 5 5 0Advanced example
bm_adv <- pc_benchmark(
ids = rep(2244, 100),
fn = function(chunk_ids, ...) {
# transport-style worker without network side effects
pc_response(
'{"IdentifierList":{"CID":[2244]}}',
request = list(domain = "compound", namespace = "cid", identifier = chunk_ids)
)
},
chunk_sizes = c(10, 25),
parallel_options = c(FALSE)
)
bm_adv
#> # A tibble: 2 × 7
#> chunk_size parallel workers elapsed_sec chunks successful_chunks failed_chunks
#> <int> <lgl> <dbl> <dbl> <int> <int> <int>
#> 1 10 FALSE 1 0.00100 10 10 0
#> 2 25 FALSE 1 0.00100 4 4 0Interpretation:
- Benchmark workers can be synthetic to isolate orchestration overhead.
Threshold Gates
Purpose:
- Scale benchmarking across scenario sizes and gate against thresholds.
Minimal example
harness_min <- pc_benchmark_harness(
fn = function(chunk_ids, ...) sum(chunk_ids),
ids = 1:60,
scenario_sizes = c(10L, 20L),
chunk_sizes = c(5L),
parallel_options = FALSE
)
harness_min$summary
#> # A tibble: 2 × 9
#> scenario_size runs min_elapsed_sec max_elapsed_sec max_failed_chunk_ratio
#> <int> <int> <dbl> <dbl> <dbl>
#> 1 10 1 0.00100 0.00100 0
#> 2 20 1 0.00100 0.00100 0
#> # ℹ 4 more variables: elapsed_threshold <dbl>, failed_ratio_threshold <dbl>,
#> # all_runs_pass <lgl>, any_run_pass <lgl>Typical example
harness_path <- file.path(tempdir(), "pubchemr-benchmark-report.md")
harness_typical <- pc_benchmark_harness(
fn = function(chunk_ids, ...) sum(chunk_ids),
ids = 1:100,
scenario_sizes = c(10L, 30L),
chunk_sizes = c(5L, 10L),
parallel_options = FALSE,
report_path = harness_path,
report_format = "markdown"
)
file.exists(harness_path)
#> [1] TRUEAdvanced example
custom_thresholds <- list(
elapsed_sec = c(`10` = 60, `30` = 120),
failed_chunk_ratio = c(`10` = 0, `30` = 0)
)
rds_report_path <- file.path(tempdir(), "pubchemr-benchmark-report.rds")
harness_adv <- pc_benchmark_harness(
fn = function(chunk_ids, ...) sum(chunk_ids),
ids = NULL,
scenario_sizes = c(10L, 30L),
chunk_sizes = c(5L),
parallel_options = FALSE,
thresholds = custom_thresholds,
id_generator = function(n) rep(2244L, n),
report_path = rds_report_path,
report_format = "rds"
)
list(
summary = harness_adv$summary %>% select(scenario_size, all_runs_pass, elapsed_threshold, failed_ratio_threshold),
report_exists = file.exists(rds_report_path),
report_class = class(readRDS(rds_report_path))[1]
)
#> $summary
#> # A tibble: 2 × 4
#> scenario_size all_runs_pass elapsed_threshold failed_ratio_threshold
#> <int> <lgl> <dbl> <dbl>
#> 1 10 TRUE 60 0
#> 2 30 TRUE 120 0
#>
#> $report_exists
#> [1] TRUE
#>
#> $report_class
#> [1] "PubChemBenchmarkReport"Interpretation:
- Harness outputs are CI-friendly and suitable for regression gating.
id_generatoris useful when you want scenario sizes to scale without recycling a short seed identifier vector.- Report artifacts can be emitted as Markdown, CSV, or RDS depending on the downstream consumer.
Analysis Layer
This section covers pc_assay_activity_long(),
pc_activity_outcome_map(),
pc_activity_matrix(), pc_cross_domain_join(),
pc_feature_table(), pc_model_matrix(),
pc_export_model_data(), pc_to_rcdk(),
pc_to_chemminer(), and
pc_lifecycle_policy().
Section contract:
- This is the section where retrieval becomes analysis.
- Inputs: normalized assay payloads, compound feature tables, and optional bridge dependencies.
- Outputs: modeling-ready long tables, matrices, joins, and exported artifacts.
- Interpret the outputs by asking whether the structure is now suitable for the next scientific task: join, plot, rank, model, or export.
- Pitfalls: inconsistent ID typing (
CID,AID,SID) and silent join-key mismatches. - Performance note: perform heavy retrieval once, persist to disk, and iterate analysis steps offline.
Built-In Fixtures
assay_payload <- pc_example_assaysummary_payload()
feature_tbl_synthetic <- pc_example_feature_table() %>%
mutate(
XLogP = c(1.2, 3.1, 2.7),
TPSA = c(63.6, 37.3, 48.4),
CanonicalSMILES = c(
"CC(=O)OC1=CC=CC=C1C(=O)O",
"CC(C)CC1=CC=C(C=C1)C(C)C(=O)O",
"CN1C=NC2=C1C(=O)N(C(=O)N2)C"
)
)
assay_long <- pc_assay_activity_long(x = assay_payload)
list(
assay_rows = nrow(assay_long),
assay_cols = names(assay_long),
feature_rows = nrow(feature_tbl_synthetic),
feature_cols = names(feature_tbl_synthetic)
)
#> $assay_rows
#> [1] 3
#>
#> $assay_cols
#> [1] "AID" "SID" "CID"
#> [4] "ActivityOutcome" "ActivityValue_uM" "ActivityOutcomeValue"
#>
#> $feature_rows
#> [1] 3
#>
#> $feature_cols
#> [1] "CID" "MolecularWeight" "XLogP"
#> [4] "TPSA" "HBondDonorCount" "HBondAcceptorCount"
#> [7] "CanonicalSMILES"Interpretation:
- These helpers are exported specifically for deterministic examples, tests, and offline workflow scaffolding.
assay_longis the canonical long format for downstream matrix/modeling operations.
Long Assay Tables
Purpose:
- Normalize assay-summary payloads to long tabular structure.
- Key inputs:
x(payload orPubChemResult) oridentifier+namespace. - Key output:
PubChemTablewith canonical columns (CID,AID,ActivityOutcome, optionalActivityOutcomeValue).
Minimal example
pc_assay_activity_long(x = assay_payload)
#> # A tibble: 3 × 6
#> AID SID CID ActivityOutcome ActivityValue_uM ActivityOutcomeValue
#> <chr> <chr> <chr> <chr> <dbl> <dbl>
#> 1 1001 9001 2244 Active 1.2 1
#> 2 1002 9002 2244 Inactive NA 0
#> 3 1003 9003 3672 Inconclusive 12.5 NATypical example
pc_assay_activity_long(
x = assay_payload,
unique_rows = TRUE,
add_outcome_value = TRUE
)
#> # A tibble: 3 × 6
#> AID SID CID ActivityOutcome ActivityValue_uM ActivityOutcomeValue
#> <chr> <chr> <chr> <chr> <dbl> <dbl>
#> 1 1001 9001 2244 Active 1.2 1
#> 2 1002 9002 2244 Inactive NA 0
#> 3 1003 9003 3672 Inconclusive 12.5 NAAdvanced example
pc_assay_activity_long(
x = assay_payload,
add_outcome_value = TRUE,
strict_outcome = FALSE,
unknown_outcome = -1
)
#> # A tibble: 3 × 6
#> AID SID CID ActivityOutcome ActivityValue_uM ActivityOutcomeValue
#> <chr> <chr> <chr> <chr> <dbl> <dbl>
#> 1 1001 9001 2244 Active 1.2 1
#> 2 1002 9002 2244 Inactive NA 0
#> 3 1003 9003 3672 Inconclusive 12.5 NAInterpretation:
ActivityOutcomeValueis derived with explicit mapping controls.
Failure-object example
assay_long_result <- pc_assay_activity_long(
x = 1,
error_mode = "result"
)
summarize_any(assay_long_result)
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemResult InvalidInput 1 4Interpretation:
error_mode = "result"is the right choice when long-table normalization is part of a larger orchestrated pipeline that should not stop immediately.
Pitfalls:
- When using live retrieval via
identifier, failed chunks now error explicitly. - Use
error_mode = "result"when you need failure objects instead of hard stops.
Outcome Encoding
Purpose:
- Convert textual assay outcomes to numeric model-ready values.
- Key inputs: outcome labels plus optional custom
map. - Key output: numeric vector aligned to input order.
Minimal example
pc_activity_outcome_map(c("Active", "Inactive", "Inconclusive"))
#> [1] 1 0 NATypical example
pc_activity_outcome_map(
c("Hit", "Non-Hit", "UnknownLabel"),
strict = FALSE,
unknown = -1
)
#> [1] 1 0 -1Advanced example
safe_call(
pc_activity_outcome_map(
c("Active", "mystery-state"),
strict = TRUE
)
)
#> $message
#> [1] "Unknown activity outcome label(s): mystery-state. Provide 'map' entries or set strict = FALSE."
#>
#> attr(,"class")
#> [1] "pc_error"Interpretation:
- Use
strict = TRUEin production scoring to fail on unexpected labels. - For exploratory analysis,
strict = FALSEwith explicitunknownmakes data-loss tradeoffs explicit.
Activity Matrices
Purpose:
- Build dense or sparse CID x AID activity matrices.
- Key inputs: long assay table with
CID,AID, and mapped numeric outcome. - Key outputs: dense tibble/matrix or
PubChemSparseActivityMatrix.
Minimal example
mat_dense <- pc_activity_matrix(assay_long)
mat_dense
#> # A tibble: 2 × 4
#> CID AID_1001 AID_1002 AID_1003
#> <chr> <dbl> <dbl> <dbl>
#> 1 2244 1 0 NA
#> 2 3672 NA NA NATypical example
mat_sparse <- pc_activity_matrix(
assay_long,
output = "sparse",
aggregate = "max"
)
mat_sparse
#>
#> PubChemSparseActivityMatrix
#>
#> - Rows (compounds): 2
#> - Columns (assays): 3
#> - Non-zero entries: 3
#> - Implicit fill: NAAdvanced example
assay_with_dupes <- bind_rows(
assay_long,
tibble(CID = "2244", AID = "1001", ActivityOutcome = "Inactive", ActivityValue_uM = 0.5, ActivityOutcomeValue = 0)
)
pc_activity_matrix(
assay_with_dupes,
aggregate = "mean",
output = "tibble",
fill = NA_real_,
prefix = "AID_"
)
#> # A tibble: 2 × 4
#> CID AID_1001 AID_1002 AID_1003
#> <chr> <dbl> <dbl> <dbl>
#> 1 2244 0.5 0 NA
#> 2 3672 NA NA NAInterpretation:
- Choose
aggregatebased on assay repeat semantics (max,mean,first). - Sparse output is preferred when assays are high-dimensional and mostly missing.
Cross-Domain Joins
Purpose:
- Join compounds, substances, assays, and targets into one analysis table.
- Key inputs: domain tables plus join keys (
bylist andjointype). - Key output: unified table for downstream feature/outcome engineering.
Minimal example
pc_cross_domain_join(
compounds = feature_tbl_synthetic,
assays = assay_long %>% select(CID, AID, ActivityOutcome)
)
#> # A tibble: 4 × 9
#> CID MolecularWeight XLogP TPSA HBondDonorCount HBondAcceptorCount
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 2244 180. 1.2 63.6 1 4
#> 2 2244 180. 1.2 63.6 1 4
#> 3 3672 206. 3.1 37.3 0 2
#> 4 5957 194. 2.7 48.4 1 4
#> # ℹ 3 more variables: CanonicalSMILES <chr>, AID <chr>, ActivityOutcome <chr>Typical example
substances_tbl <- tibble(CID = c("2244", "3672"), SID = c("111", "222"))
target_tbl <- tibble(AID = c("1001", "1002"), target_symbol = c("PTGS1", "PTGS2"))
pc_cross_domain_join(
compounds = feature_tbl_synthetic,
substances = substances_tbl,
assays = assay_long %>% select(CID, AID, ActivityOutcomeValue),
targets = target_tbl,
join = "left"
)
#> # A tibble: 4 × 11
#> CID MolecularWeight XLogP TPSA HBondDonorCount HBondAcceptorCount
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 2244 180. 1.2 63.6 1 4
#> 2 2244 180. 1.2 63.6 1 4
#> 3 3672 206. 3.1 37.3 0 2
#> 4 5957 194. 2.7 48.4 1 4
#> # ℹ 5 more variables: CanonicalSMILES <chr>, SID <chr>, AID <chr>,
#> # ActivityOutcomeValue <dbl>, target_symbol <chr>Advanced example
pc_cross_domain_join(
compounds = feature_tbl_synthetic,
assays = assay_long %>% select(CID, AID, ActivityOutcomeValue),
by = list(compound_assay = "CID"),
join = "full"
)
#> # A tibble: 4 × 9
#> CID MolecularWeight XLogP TPSA HBondDonorCount HBondAcceptorCount
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 2244 180. 1.2 63.6 1 4
#> 2 2244 180. 1.2 63.6 1 4
#> 3 3672 206. 3.1 37.3 0 2
#> 4 5957 194. 2.7 48.4 1 4
#> # ℹ 3 more variables: CanonicalSMILES <chr>, AID <chr>,
#> # ActivityOutcomeValue <dbl>Interpretation:
- Explicit join-key maps avoid silent mismatches in heterogeneous datasets.
- Standardize ID types (
character) before joining to prevent accidental row loss.
Feature Tables
Purpose:
- Retrieve modeling-ready property tables from PubChem.
- Key inputs: identifiers, property set, namespace, numeric coercion policy.
- Key output: feature tibble with transport metadata removed.
Minimal example
feature_try <- safe_call(
pc_feature_table(
identifier = c(2244, 3672),
properties = c("MolecularWeight", "XLogP", "TPSA"),
namespace = "cid",
offline = TRUE
)
)
summarize_any(feature_try)
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE pc_error Property retrieval failed: Offline mode enabled an… NA NATypical example
if (run_live) {
feature_live <- pc_feature_table(
identifier = c(2244, 3672),
properties = c("MolecularWeight", "XLogP", "TPSA"),
namespace = "cid",
numeric_only = TRUE,
cache = TRUE
)
feature_live
} else {
feature_tbl_synthetic
}
#> # A tibble: 3 × 7
#> CID MolecularWeight XLogP TPSA HBondDonorCount HBondAcceptorCount
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 2244 180. 1.2 63.6 1 4
#> 2 3672 206. 3.1 37.3 0 2
#> 3 5957 194. 2.7 48.4 1 4
#> # ℹ 1 more variable: CanonicalSMILES <chr>Advanced example
if (run_live) {
feature_live_adv <- pc_feature_table(
identifier = c(2244, 3672, 5957),
properties = c("MolecularWeight", "XLogP", "TPSA", "HBondDonorCount", "HBondAcceptorCount"),
namespace = "cid",
numeric_only = TRUE,
cache = TRUE,
force_refresh = TRUE
)
feature_live_adv
} else {
feature_result_adv <- pc_feature_table(
identifier = c(2244, 3672, 5957),
properties = c("MolecularWeight", "XLogP", "TPSA", "HBondDonorCount", "HBondAcceptorCount"),
namespace = "cid",
offline = TRUE,
error_mode = "result"
)
summarize_any(feature_result_adv)
}
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemRecord OfflineCacheMiss 1 4Interpretation:
- In offline mode,
pc_feature_table()intentionally fails on cache miss. - In live mode, it is the preferred feature-ingestion entry point.
- Use
error_mode = "result"when pipeline orchestration requires typed failures.
Model Matrices
Purpose:
- Convert mixed feature tables to numeric predictor matrix + optional outcome.
- Key inputs: tabular data,
outcome, ID columns, NA and scaling policy. - Key output:
PubChemModelMatrixwithx, optionaly, and ID metadata.
Minimal example
mm_min <- pc_model_matrix(
x = assay_long %>%
select(CID, AID, ActivityOutcomeValue) %>%
mutate(dummy = as.numeric(factor(AID))),
outcome = "ActivityOutcomeValue",
id_cols = c("CID", "AID")
)
mm_min
#>
#> PubChemModelMatrix
#>
#> - Rows: 3
#> - Features: 1
#> - Outcome: ProvidedTypical example
joined_tbl <- pc_cross_domain_join(
compounds = feature_tbl_synthetic,
assays = assay_long %>% select(CID, AID, ActivityOutcomeValue)
)
mm_typical <- pc_model_matrix(
x = joined_tbl,
outcome = "ActivityOutcomeValue",
id_cols = c("CID", "AID"),
na_fill = 0,
scale = TRUE
)
dim(mm_typical$x)
#> [1] 4 5Advanced example
mm_adv <- pc_model_matrix(
x = joined_tbl,
outcome = NULL,
id_cols = c("CID", "AID", "ActivityOutcomeValue"),
na_fill = 0,
scale = FALSE
)
list(features = length(mm_adv$feature_names), has_outcome = !is.null(mm_adv$y), has_ids = !is.null(mm_adv$ids))
#> $features
#> [1] 5
#>
#> $has_outcome
#> [1] FALSE
#>
#> $has_ids
#> [1] TRUEInterpretation:
pc_model_matrix()is strict about non-empty numeric predictor sets.- Inspect
feature_namesbefore modeling to ensure expected predictors survived preprocessing.
Data Export
Purpose:
- Persist model-ready data bundles to CSV/RDS.
- Key inputs:
PubChemModelMatrixor data frame, output path, format flags. - Key output: invisible metadata list (
path,format,rows,cols).
Minimal example
out_csv <- file.path(tempdir(), "pubchemr_mm_min.csv")
meta_csv <- pc_export_model_data(mm_typical, path = out_csv, format = "csv")
meta_csv
#> $path
#> [1] "/private/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T/RtmpjLjM6G/pubchemr_mm_min.csv"
#>
#> $format
#> [1] "csv"
#>
#> $rows
#> [1] 4
#>
#> $cols
#> [1] 8Typical example
out_rds <- file.path(tempdir(), "pubchemr_mm_typical.rds")
meta_rds <- pc_export_model_data(mm_typical, path = out_rds, format = "rds")
meta_rds
#> $path
#> [1] "/private/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T/RtmpjLjM6G/pubchemr_mm_typical.rds"
#>
#> $format
#> [1] "rds"
#>
#> $rows
#> [1] 4
#>
#> $cols
#> [1] 8Advanced example
out_csv_no_ids <- file.path(tempdir(), "pubchemr_mm_no_ids.csv")
meta_csv_no_ids <- pc_export_model_data(
mm_typical,
path = out_csv_no_ids,
format = "csv",
include_ids = FALSE,
include_outcome = FALSE
)
meta_csv_no_ids
#> $path
#> [1] "/private/var/folders/dj/y28dp44x303ggfg6rg8n2v0h0000gn/T/RtmpjLjM6G/pubchemr_mm_no_ids.csv"
#>
#> $format
#> [1] "csv"
#>
#> $rows
#> [1] 4
#>
#> $cols
#> [1] 5Interpretation:
- Keep export settings explicit to guarantee schema stability across runs.
- Use
tempdir()in reproducible examples; in projects, write to explicit versioned artifact directories.
Similarity Search
Purpose:
- Perform similarity-driven ID retrieval with configurable search strategy.
- Key inputs: structure identifier, namespace, threshold, target
namespace (
to). - Key output: typed ID-mapping style result convertible with
as_tibble()orretrieve().
Minimal example
sim_min <- safe_call(
pc_similarity_search(
identifier = "CC(=O)OC1=CC=CC=C1C(=O)O",
namespace = "smiles",
offline = TRUE
)
)
summarize_any(sim_min)
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemSimilarityResult OfflineCacheMiss 1 5Typical example
if (run_live) {
sim_live <- pc_similarity_search(
identifier = "CC(=O)OC1=CC=CC=C1C(=O)O",
namespace = "smiles",
threshold = 90,
max_records = 50,
cache = TRUE
)
as_tibble(sim_live) %>% head(10)
} else {
tibble(note = "Skipped live similarity retrieval.")
}
#> # A tibble: 1 × 1
#> note
#> <chr>
#> 1 Skipped live similarity retrieval.Advanced example
sim_adv <- pc_similarity_search(
identifier = "CCO",
namespace = "smiles",
searchtype = "fastsimilarity_2d",
threshold = 80,
to = "aids",
max_records = 100,
offline = TRUE
)
list(
summary = summarize_any(sim_adv),
request = list(
searchtype = request_args(sim_adv, "searchtype"),
to = request_args(sim_adv, "to"),
threshold = request_args(sim_adv, "options")$Threshold,
max_records = request_args(sim_adv, "options")$MaxRecords
)
)
#> $summary
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemSimilarityResult OfflineCacheMiss 1 5
#>
#> $request
#> $request$searchtype
#> [1] "fastsimilarity_2d"
#>
#> $request$to
#> [1] "aids"
#>
#> $request$threshold
#> [1] 80
#>
#> $request$max_records
#> [1] 100Interpretation:
- Use
searchtype,threshold, andtointentionally; these strongly affect downstream payload shape. - Similarity search can trigger listkey polling internally and
optionally remap resulting CIDs to
SID/AIDtargets before returning. - Lower thresholds expand recall but can substantially increase downstream join volume.
Bioactivity Retrieval
Purpose:
- Retrieve high-dimensional bioactivity records from the SDQ endpoint.
- Key inputs: identifier/namespace resolution, collection, limit/order, and transport policy.
- Key output:
PubChemTablewith SDQ-specific columns (variable by record).
Minimal example
if (run_live) {
sdq_min <- pc_sdq_bioactivity(2244, namespace = "cid", limit = 100L, rate_limit = FALSE)
summarize_any(sdq_min)
} else {
tibble(note = "Skipped (live-only function).")
}
#> # A tibble: 1 × 1
#> note
#> <chr>
#> 1 Skipped (live-only function).Typical example
if (run_live) {
sdq_typical <- pc_sdq_bioactivity(
identifier = 2244,
namespace = "cid",
collection = "bioactivity",
limit = 200L,
order = "activity,asc",
cache = TRUE,
cache_dir = work_cache,
cache_ttl = 3600,
rate_limit = FALSE
)
sdq_typical %>% head(5)
} else {
tibble(note = "Skipped (live-only function).")
}
#> # A tibble: 1 × 1
#> note
#> <chr>
#> 1 Skipped (live-only function).Advanced example
if (run_live) {
sdq_adv <- pc_sdq_bioactivity(
identifier = "aspirin",
namespace = "name",
limit = 250L,
cache = TRUE,
cache_dir = work_cache,
force_refresh = TRUE,
rate_limit = FALSE
)
summarize_any(sdq_adv)
} else {
sdq_adv <- pc_sdq_bioactivity(
identifier = "aspirin",
namespace = "name",
cache = TRUE,
cache_dir = work_cache,
offline = TRUE,
error_mode = "result"
)
list(
summary = summarize_any(sdq_adv),
request = request_args(sdq_adv)[c("identifier", "namespace", "caller")]
)
}
#> $summary
#> # A tibble: 1 × 5
#> ok class note rows cols
#> <lgl> <chr> <chr> <int> <int>
#> 1 FALSE PubChemResult OfflineCacheMiss 1 4
#>
#> $request
#> $request$identifier
#> [1] "aspirin"
#>
#> $request$namespace
#> [1] "name"
#>
#> $request$<NA>
#> NULLInterpretation:
- SDQ can return many columns; always inspect schema before modeling.
error_mode = "result"is recommended for robust pipelines that must continue after some retrieval failures.
Cheminformatics Bridges
Purpose:
- Bridge PubChem feature tables to optional cheminformatics ecosystems.
- Key inputs: feature table plus valid SMILES column (and optional ID column).
- Key outputs:
rcdkmolecule lists orChemmineRSDF objects.
Minimal examples
rcdk_min <- safe_call(pc_to_rcdk(feature_tbl_synthetic, smiles_col = "CanonicalSMILES", id_col = "CID"))
chemminer_min <- safe_call(pc_to_chemminer(feature_tbl_synthetic, smiles_col = "CanonicalSMILES"))
bind_rows(
list(pc_to_rcdk = summarize_any(rcdk_min), pc_to_chemminer = summarize_any(chemminer_min)),
.id = "function"
)
#> # A tibble: 2 × 6
#> `function` ok class note rows cols
#> <chr> <lgl> <chr> <chr> <int> <int>
#> 1 pc_to_rcdk FALSE pc_error Package 'rcdk' is required for pc_… NA NA
#> 2 pc_to_chemminer FALSE pc_error Package 'ChemmineR' is required fo… NA NATypical examples
if (requireNamespace("rcdk", quietly = TRUE)) {
mols <- pc_to_rcdk(feature_tbl_synthetic, smiles_col = "CanonicalSMILES", id_col = "CID")
length(mols)
} else {
"rcdk not installed; conversion skipped."
}
#> [1] "rcdk not installed; conversion skipped."
if (requireNamespace("ChemmineR", quietly = TRUE)) {
sdf_obj <- pc_to_chemminer(feature_tbl_synthetic, smiles_col = "CanonicalSMILES")
class(sdf_obj)
} else {
"ChemmineR not installed; conversion skipped."
}
#> [1] "ChemmineR not installed; conversion skipped."Advanced examples
if (requireNamespace("rcdk", quietly = TRUE)) {
mols_named <- pc_to_rcdk(feature_tbl_synthetic, smiles_col = "CanonicalSMILES", id_col = "CID")
names(mols_named)
} else {
"rcdk not installed; advanced conversion skipped."
}
#> [1] "rcdk not installed; advanced conversion skipped."
if (requireNamespace("ChemmineR", quietly = TRUE)) {
sdf_adv <- pc_to_chemminer(feature_tbl_synthetic, smiles_col = "CanonicalSMILES")
length(sdf_adv)
} else {
"ChemmineR not installed; advanced conversion skipped."
}
#> [1] "ChemmineR not installed; advanced conversion skipped."Interpretation:
- These are optional bridges; they should be feature-gated in production code.
- Run
pc_capabilities()early to detect optional dependency availability.
Lifecycle Policy
Purpose:
- Make compatibility/deprecation policy explicit in scripts and governance docs.
- Key input: none (policy is package-defined metadata).
- Key output: policy tibble for programmatic checks.
Minimal example
pc_lifecycle_policy()
#> # A tibble: 2 × 5
#> stream stability support_window deprecation_notice breaking_change_window
#> <chr> <chr> <chr> <chr> <chr>
#> 1 legacy maintenance bugfix-only >= 1 minor release major release only
#> 2 nextgen stable minor+patch >= 2 minor releases major release onlyTypical example
pc_lifecycle_policy() %>% filter(stream == "nextgen")
#> # A tibble: 1 × 5
#> stream stability support_window deprecation_notice breaking_change_window
#> <chr> <chr> <chr> <chr> <chr>
#> 1 nextgen stable minor+patch >= 2 minor releases major release onlyAdvanced example
policy <- pc_lifecycle_policy()
stopifnot(any(policy$stream == "nextgen"), any(policy$stream == "legacy"))
policy
#> # A tibble: 2 × 5
#> stream stability support_window deprecation_notice breaking_change_window
#> <chr> <chr> <chr> <chr> <chr>
#> 1 legacy maintenance bugfix-only >= 1 minor release major release only
#> 2 nextgen stable minor+patch >= 2 minor releases major release onlyInterpretation:
- Treat lifecycle policy as operational metadata, not only documentation.
Integrated Pipelines
This section provides two coherent pipelines users can adapt directly.
Section contract:
- Read this section when you want end-to-end templates rather than individual function examples.
- Inputs: deterministic fixtures, controlled cache paths, and local worker functions.
- Outputs: complete, portable workflow skeletons for analysis and orchestration.
- Interpret the outputs by checking whether every stage produces an object that is ready for the next stage without manual repair.
- Pitfalls: introducing live retrieval before pipeline logic is validated.
- Performance note: benchmark with local deterministic workers first, then replace worker internals with network-bound retrieval.
Pipeline A
Goal:
- Build model-ready artifacts without requiring internet access.
# 1) Start from assay payload normalization.
assay_long_a <- pc_assay_activity_long(x = assay_payload, add_outcome_value = TRUE)
# 2) Build dense and sparse activity matrices.
activity_dense_a <- pc_activity_matrix(assay_long_a, output = "tibble")
activity_sparse_a <- pc_activity_matrix(assay_long_a, output = "sparse")
# 3) Join synthetic features with assay outcomes.
joined_a <- pc_cross_domain_join(
compounds = feature_tbl_synthetic,
assays = assay_long_a %>% select(CID, AID, ActivityOutcomeValue)
)
# 4) Create model matrix.
mm_a <- pc_model_matrix(
x = joined_a,
outcome = "ActivityOutcomeValue",
id_cols = c("CID", "AID"),
na_fill = 0,
scale = TRUE
)
# 5) Export artifacts.
out_a_csv <- file.path(tempdir(), "pipeline_a_model.csv")
out_a_rds <- file.path(tempdir(), "pipeline_a_model.rds")
meta_a_csv <- pc_export_model_data(mm_a, path = out_a_csv, format = "csv")
meta_a_rds <- pc_export_model_data(mm_a, path = out_a_rds, format = "rds")
list(
assay_rows = nrow(assay_long_a),
dense_dim = dim(activity_dense_a),
sparse_dim = dim(activity_sparse_a$x),
model_dim = dim(mm_a$x),
csv_exists = file.exists(meta_a_csv$path),
rds_exists = file.exists(meta_a_rds$path)
)
#> $assay_rows
#> [1] 3
#>
#> $dense_dim
#> [1] 2 4
#>
#> $sparse_dim
#> [1] 2 3
#>
#> $model_dim
#> [1] 4 5
#>
#> $csv_exists
#> [1] TRUE
#>
#> $rds_exists
#> [1] TRUEInterpretation:
- This pipeline is fully runnable in a clean offline session.
- It demonstrates a complete nextgen modeling path from assay data to export.
Pipeline B
Goal:
- Demonstrate checkpointed orchestration and threshold-gated benchmarking.
pipe_b_cp_dir <- file.path(tempdir(), "pipeline_b_checkpoints")
pipe_b_cp_id <- "pipeline-b-demo"
# Chunked workflow using a deterministic local worker.
pipe_b_batch <- pc_batch(
ids = 1:24,
fn = function(chunk_ids, ...) {
tibble(id = chunk_ids, value = chunk_ids * 2, parity = ifelse(chunk_ids %% 2 == 0, "even", "odd"))
},
chunk_size = 6,
checkpoint_dir = pipe_b_cp_dir,
checkpoint_id = pipe_b_cp_id
)
# Resume using the same checkpoint.
pipe_b_resume <- pc_resume_batch(
fn = function(chunk_ids, ...) tibble(id = chunk_ids, value = chunk_ids * 2),
checkpoint_dir = pipe_b_cp_dir,
checkpoint_id = pipe_b_cp_id
)
# Benchmark and harness on local worker.
pipe_b_bm <- pc_benchmark(
ids = 1:120,
fn = function(chunk_ids, ...) sum(chunk_ids),
chunk_sizes = c(10, 20, 40),
parallel_options = FALSE
)
pipe_b_harness <- pc_benchmark_harness(
fn = function(chunk_ids, ...) sum(chunk_ids),
ids = 1:240,
scenario_sizes = c(20L, 60L),
chunk_sizes = c(10L, 20L),
parallel_options = FALSE,
report_path = file.path(tempdir(), "pipeline_b_harness.md"),
report_format = "markdown"
)
list(
batch_chunks = length(pipe_b_batch$chunks),
resumed = pipe_b_resume$checkpoint$resumed,
benchmark_rows = nrow(pipe_b_bm),
harness_rows = nrow(pipe_b_harness$summary)
)
#> $batch_chunks
#> [1] 4
#>
#> $resumed
#> [1] TRUE
#>
#> $benchmark_rows
#> [1] 3
#>
#> $harness_rows
#> [1] 2Interpretation:
- This pipeline is ideal for stress-testing orchestration logic before live runs.
- It combines
pc_batch(),pc_resume_batch(),pc_benchmark(), andpc_benchmark_harness().
Troubleshooting
Section contract:
- Read this section whenever a workflow returns the right object type
but the wrong state, especially
success = FALSE. - Inputs: failed results, cache diagnostics, and execution metadata.
- Outputs: reproducible diagnostics and corrective actions.
- Interpret failures by reading typed fields first
(
error$code,error$message,pending,from_cache) before changing the workflow. - Pitfalls: debugging only printed errors instead of typed fields.
Offline Cache Misses
Cause:
offline = TRUEwith no cached response for that request.
Diagnostic:
x <- pc_request(identifier = 2244, cache = TRUE, offline = TRUE)
tibble(success = x$success, error_code = x$error$code, error_message = x$error$message)
#> # A tibble: 1 × 3
#> success error_code error_message
#> <lgl> <chr> <chr>
#> 1 FALSE OfflineCacheMiss Offline mode enabled and no cached response found fo…Best practice:
- Run once online with
cache = TRUE, then replay withoffline = TRUE.
Wrapper Failures
Diagnostic pattern:
x <- safe_call(pc_property(2244, properties = "MolecularWeight", offline = TRUE))
if (inherits(x, "pc_error")) {
x$message
} else {
list(success = x$success, error = if (x$success) NA_character_ else x$error$message)
}
#> $success
#> [1] FALSE
#>
#> $error
#> [1] "Offline mode enabled and no cached response found for this request."Best practice:
- Branch on
successand inspecterror$code/error$message. - Use
retrieve(x, "error/code", .to.data.frame = FALSE)when building programmatic routing logic.
Offline Feature Tables
Cause:
- It is a higher-level wrapper that expects successful property retrieval.
Diagnostic:
safe_call(
pc_feature_table(
identifier = c(2244, 3672),
properties = c("MolecularWeight", "XLogP"),
offline = TRUE
)
)
#> $message
#> [1] "Property retrieval failed: Offline mode enabled and no cached response found for this request."
#>
#> attr(,"class")
#> [1] "pc_error"Best practice:
- Use
error_mode = "result"for typed failures in orchestration code. - Use live mode (optionally with cache) for feature ingestion, then persist and replay locally.
Intermittent Chunk Failures
Best practice checklist:
- Lower
chunk_size. - Increase retries/timeouts via
pc_config(). - Enable checkpointing and use
pc_resume_batch(). - Track error codes per chunk from
PubChemBatchResult$error.
Reproducibility
Checklist:
- Set seed at session start (
set.seed(...)) for local stochastic steps. - Pin transport config with
pc_profile()+ explicitpc_config()overrides. - Use
tempdir()for vignette/testing artifacts. - Keep live-network examples guarded (
run_live).
Final recommendations
For production nextgen workflows:
- Start with
pc_profile()and explicitpc_config()overrides. - Build all control flow around typed contracts (
success,error, metadata). - Use checkpointed batching for recoverability.
- Gate performance with benchmark harness thresholds.
- Separate data retrieval (possibly live + cache) from offline modeling stages.
- Use
retrieve()(oras_tibble()) consistently to normalizepc_*outputs. - Keep optional cheminformatics bridges dependency-guarded.